FTP area updated
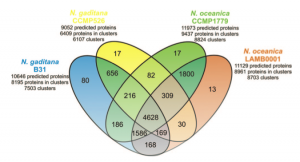
![]() We updated our FTP area, now including datasets from other Nannochloropsis species and strains and the list of families of orthologous proteins obtained from the comparison of N. gaditana and N. oceeanica predicted proteins.
We updated our FTP area, now including datasets from other Nannochloropsis species and strains and the list of families of orthologous proteins obtained from the comparison of N. gaditana and N. oceeanica predicted proteins.
Families of orthologous proteins: file type
N.gaditanaB-31|Naga_100019g47.1 N.gaditanaCCMP526|Nga20827 nudix hydrolase;
Even though the differences and the imprecisions of the various gene predictions probably play a major role in the determination of the differences among the two species, a close look at the lists of proteins that are putatively assigned as characteristic of each species my reserve interesting surprises!
References
- Corteggiani Carpinelli, E. et al. “Chromosome scale genome assembly and transcriptome profiling of Nannochloropsis gaditana in nitrogen depletion.” Molecular Plant (2014) 7 (2): 323-335.doi: 10.1093/mp/sst120
We’re looking for new authors and contributors
 Are you an expert about Nannochloropsis? Is your research focused on this organism?
Are you an expert about Nannochloropsis? Is your research focused on this organism?
Consider becoming a new author of this blog and contributing to the scientific discussion!
Why don’t you start with advertising your recent publication or commenting on the posts of this blog?
Is there any hot topic that you would like to discuss or any suggestion that you would like to ask to this scientific community?
Fill this form and we will be happy to consider you application as new author

Recent Comments