Determining Nannochloropsis cell wall and more
 Nannochloropsis’ whole cells were recently analysed using 13C-SS-NMR spectroscopy (reference n 1). Cells were grown autotrophically in a sodium [13C] bicarbonate environment and harvested during the exponential phase.
Nannochloropsis’ whole cells were recently analysed using 13C-SS-NMR spectroscopy (reference n 1). Cells were grown autotrophically in a sodium [13C] bicarbonate environment and harvested during the exponential phase.
Quantitative spectra of 13C labelled cells of Nannochloropsis oculata were analysed for resonance picks which are characteristic of carbohydrates and lipids, thus providing, among other interesting observations, useful insights on the cell wall composition of the microalga.
Carbohydrate resonances in microalgae are expected from the cell walls, the headgroup of galactolipids and the storage polysaccharides.
Cell wall
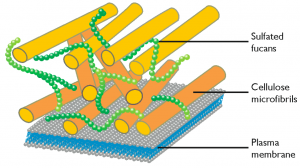
Analysis of the RINEPT and CP spectra of N. oculata allowed to detect resonances in which chemical shifts agreed with those reported for cellulose. The cellulose anomeric carbon C1 only appeared in the rigid (CP) spectrum, the remaining carbons showed up in the spectra of both rigid and mobile zones. The cellulose was attributed to the cell wall (reference n 1). Analysis of the genes present in Nannochloropsis genome also showed the presence of a full set of genes that can be attributed to pathways leading to synthesis and modification of cellulose (reference n 2).
Storage sugars
Chemical shifts corresponding to both β-(1 → 3) and β-(1 → 6) glycans were detected and assigned to chrysolaminarin. Also in this case some of the resonances were exclusively detected in the CP spectra but the remaining resonances appeared on both rigid and mobile zones. The relative mobility observed could be compatible with the accumulation of chrysolaminarins in solution inside the vacuoles (reference n 1). In this case again genes associated to the biosynthesis of chrysolaminarins are found in the genome (reference n 2).
Lipids
Quantitative and qualitative analysis of the lipid content of various Nannochloropsis cultures based on MASS-spectrometry were previously reported (references n 3 and n 4). In this recent work by Arnold and coworkers (reference n 1) lipid signals were analysed from 13C-SS-NMR spectra of whole cells grown photoautotrophically.
The peak assignment showed the presence of galactosyldiacylglycerols. Peaks characteristic of the sulfolipid sulfoquinovosyl-diacylglycerol (SQDG) sugar head group seemed present on both the rigid (CP) and dynamic (RINEPT) spectra. The sugar moieties were better evidenced on the RINEPT spectrum while the glycerol moieties appeared better in the CP spectrum. Phosphatidylcholine (PC) was the dominant phospholipid detected and assigned to the plasma membrane.
The CP and RINEPT spectra were dominated by lipids’ methylene signal (30–33 ppm) consistent with a high content in saturated and mono-unsaturated fatty acid chains. The RINEPT spectrum also showed a strong signal at 128–130 ppm consistent with a high proportion of PUFAs, and possibly to EPA from the vacuolar and mobile TAGs.
For further reading on the subject I suggest of course to read the paper
References
- Arnold AA, Genard B, Zito F, Tremblay R, Warschawski DE, Marcotte I. “Identification of lipid and saccharide constituents of whole microalgal cells by 13C solid-state NMR.” Biochimica et Biophysica Acta (BBA) – Biomembranes. (2014) doi:10.1016/j.bbamem.2014.07.017
- Corteggiani Carpinelli E, Telatin A, Vitulo N, Forcato C, D’Angelo M, Schiavon R, Vezzi A, Giacometti GM, Morosinotto T, Valle G. “Chromosome scale genome assembly and transcriptome profiling of Nannochloropsis gaditana in nitrogen depletion.” Molecular Plant (2014) 7 (2): 323-335. doi: 10.1093/mp/sst120.
- Li J, Han D, Wang D, Ning K, Jia J, Wei L, Jing X, Huang S, Chen J, Li Y, Hu Q, Xu J “Choreography of Transcriptomes and Lipidomes of Nannochloropsis Reveals the Mechanisms of Oil Synthesis in Microalgae.” The Plant Cell April 2014. doi: http://dx.doi.org/10.1105/tpc.113.121418
- Simionato D, Block MA, La Rocca N, Jouhet J, Maréchal E, Finazzi G, Morosinotto T. (2013) The response of Nannochloropsis gaditana to nitrogen starvation includes de novo biosynthesis of triacylglycerols, a decrease of chloroplast galactolipids, and reorganisation of the photosynthetic apparatus. Eukaryotic Cell May 2013 vol. 12 no. 5 665-676
Recent Comments