FTP area updated
![]() We updated our FTP area, now including datasets from other Nannochloropsis species and strains and the list of families of orthologous proteins obtained from the comparison of N. gaditana and N. oceeanica predicted proteins.
We updated our FTP area, now including datasets from other Nannochloropsis species and strains and the list of families of orthologous proteins obtained from the comparison of N. gaditana and N. oceeanica predicted proteins.
Families of orthologous proteins: file type
N.gaditanaB-31|Naga_100019g47.1 N.gaditanaCCMP526|Nga20827 nudix hydrolase;
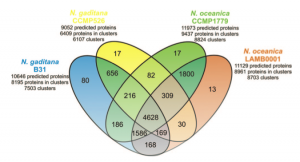
Even though the differences and the imprecisions of the various gene predictions probably play a major role in the determination of the differences among the two species, a close look at the lists of proteins that are putatively assigned as characteristic of each species my reserve interesting surprises!
References
- Corteggiani Carpinelli, E. et al. “Chromosome scale genome assembly and transcriptome profiling of Nannochloropsis gaditana in nitrogen depletion.” Molecular Plant (2014) 7 (2): 323-335.doi: 10.1093/mp/sst120
Thank you!
We currently have more than 1.000 unique visitors per month, accounting for roughly 18,000 pageviews per month. Visitors are spread across the four continents… Thank you for keeping alive Nannochloropsis.org. We are doing our best to update this site and we are also preparing new resources (to be released soon).

This week’s visits
Annotation file available
Upon user request we uploaded the GFF version of Nannochloropsis gaditana annotation. This file is in standard GFF3 format, and features refers to the assembled genome (i.e. scaffolds).
You can download the file from the Nannochloropsis genome FTP area.
Paper online
Our paper “Chromosome scale genome assembly and transcriptome profiling of Nannochloropsis gaditana in nitrogen depletion” is online.
You can access it from the Publisher’s website.
Open day
When first published, nannochloropsis.org, has been a private website providing access only upon request. We are glad to offer online the first release of the genome portal! Stay tuned (more features are coming 
Recent Comments